Serial.c File Reference
Utilities that can be different between serial & parallel codes. More...
#include "serial.h"
Go to the source code of this file.
Defines | |
| #define | ERRCODE 2 |
| Write floating point data for spatial (map) output netCDF file (new v2.8.2). | |
| #define | ERR(e) {printf("\nError in writing netCDF output map: %s\n", nc_strerror(e)); exit(ERRCODE);} |
| #define | NDIMS 3 |
Functions | |
| byte | readMap (char *filename, void *data) |
| Read data for spatial (map) input binary file. | |
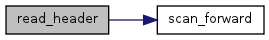
| int | read_header (char *filename) |
| Read header info for spatial (map) input binary file. | |
| void | writeMap (char *filename, void *data, int bSize, unsigned char type, int index) |
| Write data for spatial (map) output binary file. | |
| void | writeFloatMap (char *filename, char *VARNAME, float *data, int bSize, unsigned char type, int index, char *thisvarUnits, int outstep) |
| Write floating point data for spatial (map) output binary file (new v2.8.2). | |
| void | writeFloatNetCDF (char *filename, char *VARNAME, float *data, int bSize, unsigned char type, int index, char *thisvarUnits, int outstep) |
| void | write_header (char *mapFileName, int size) |
| Write header info for spatial (map) output binary file. | |
| float * | readSeriesCol (char *filename, int format, int index, int *Npt, float *TStep, int col) |
| (Unused). Read time series input data for spatial interpolations | |
| float * | get_hab_parm (char *s_parm_name, int s_parm_relval, char *parmName) |
| Read the input data file to get a habitat-specfic parameter array. | |
| char * | match_Sparm (char *s_parm_name, int s_parm_relval, char *parmName) |
| Determine if the parameter is being varied for a sensitivity analysis. | |
| float | get_global_parm (char *s_parm_name, int s_parm_relval, char *parmName) |
| Read the input data file to get a global parameter. | |
| float | get_modexperim_parm (char *s_parm_name, int s_parm_relval, char *parmName) |
| Read the input data file to get a model experiment parameter. | |
| void | open_point_lists (SeriesParm *pSeries, int nSeries) |
| Point time series output: open files and print headers. | |
| void | send_point_lists2 (SeriesParm *pSeries, int nSeries) |
| Point time series output: print time series data to file(s). | |
| void | writeSeries (void *fValue, char *label, char *desc, int N0, int N1, byte Mtype, byte format) |
| Write to a spatial data window in a debug file. | |
| void | Combine (float *fValue, char *label, int nComp, int cType, int step) |
| A variety of stat summaries in a selected spatial window (debug-related). | |
| int | getCombineIndex (char *name, int step, int type, int *last) |
| A variety of stat summaries in a selected spatial window (debug-related). | |
| void | open_debug_outFile (int index) |
| Open debug-related output files. | |
| int | init_config_file (FILE *vpFile, char term1, char term2, char term3, char term4) |
| Get format definition of output configuration file (does nothing effective!). | |
| int | skip_cnfg_space (FILE *vpFile, char *tch) |
| Skip spaces in reading (model outlist) configuration file. | |
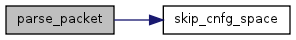
| int | parse_packet (FILE *vpFile, int *nArgs, char *test) |
| Parse through a configuration fileline to obtain the configuration commands. | |
| int | get_number (FILE *infile) |
| Get a numeric digit from a file. | |
| int | goto_index (FILE *infile, char tchar, int index) |
| Go to a particular indexed/demarked point in the habitat-specfic parameter file. | |
| float | get_Nth_parm (FILE *infile, int pIndex, int *end, int hIndex) |
| Get the N'th parameter in the habitat-specfic parameter file. | |
| float | SMDRAND (float fminVal, float fmaxVal) |
| Psuedo-random number generator (unused). | |
| void | local_setup (int argc, char **argv) |
| Does very little in serial implementation (opens a low-level debug file). | |
| void | exparam (struct nodenv *envInfo) |
| Parallel code: effectively unused in serial implementation. | |
| int | exgridinit (int dim, int *nprocs) |
| Parallel code: does nothing in serial implementation). | |
| void | exgridsplit (int nprocs, int ndim, int nprocs2[2]) |
| Parallel code: effectively unused in serial implementation. | |
| void | exgridcoord (int pnum, int rnum[2]) |
| Parallel code: effectively unused in serial implementation. | |
| void | exgridsize (int pnum, int gsize[2], int lsize[2], int lstart[2]) |
| Parallel code: effectively unused in serial implementation. | |
| void | set_async_mode (FILE *file) |
| Parallel code: does nothing in serial implementation). | |
| void | fmulti (FILE *file) |
| Parallel code: does nothing in serial implementation). | |
| void | fsingl (FILE *file) |
| Parallel code: does nothing in serial implementation). | |
| void | fasync (FILE *file) |
| Parallel code: does nothing in serial implementation). | |
| void | exchange_borders (UCHAR *map, int size) |
| Parallel code: does nothing in serial implementation). | |
| int | on_this_proc (int x, int y) |
| Parallel code: does nothing in serial implementation). | |
| void | Cplot (VOIDP Map, unsigned char Mtype, float max_value, float min_value) |
| Parallel code: does nothing in serial implementation). | |
| void | broadcastMsg (UCHAR *msgPtr) |
| Parallel code: does nothing in serial implementation). | |
| void | broadcastInt (int *iValPtr) |
| Parallel code: does nothing in serial implementation). | |
| void | broadcastChar (UCHAR *cPtr) |
| Parallel code: does nothing in serial implementation). | |
| void | broadcastData (void *dataPtr, int *dataSize) |
| Parallel code: does nothing in serial implementation). | |
| void | sync_processors () |
| Parallel code: does nothing in serial implementation). | |
| void | broadcastFloat (void *dataPtr) |
| Parallel code: does nothing in serial implementation). | |
Detailed Description
Utilities that can be different between serial & parallel codes.
This source file contains functions that are (or can be) different between single-processor (serial) vs multi-processor implementations. These functions are used in "utility" tasks that generally relate to I/O, set-up of the simulation environment, etc (and are generally called from Driver_Utilities.c).
Early versions of ELM had an analagous "Parallel.c", Express.c, etc. for code that required parallel-specfic routines. (Note: in ELM v1.0, the parallel coding efforts were merged w/ serial code (elsewhere), allowing completely generic compilation. Complexities due to vector-raster integration of canals were solved, and performance was very good (up to ~8 procs). However, that code development was halted in ~2000, and we do not anticipate parallel implementations of ELM in the near-term (Dec 2004).
Note: documented with Doxygen, which expects specific syntax within special comments.
The Everglades Landscape Model (ELM).
last updated: June 2008
Definition in file Serial.c.
Define Documentation
| #define ERRCODE 2 |
Write floating point data for spatial (map) output netCDF file (new v2.8.2).
- Parameters:
-
filename Filename (path and variable names) of data VARNAME The variable name of data data Data array of the variable bSize Byte size of data type Type, or format, of (binary) data index Index used in output sequence thisvarUnits Units of the output variable
- Remarks:
- Important: the row order of this output is reversed from all other ELM arrays. The origin of the netCDF arrays is at lower left, not the standard uppper left. Also, we converted the ELM horizontal datum from NAD27 to NAD83, only in this netCDF output. These were done for compatibility with other South Florida modeling projects that make use of the ELM output.
| #define ERR | ( | e | ) | {printf("\nError in writing netCDF output map: %s\n", nc_strerror(e)); exit(ERRCODE);} |
| #define NDIMS 3 |
Function Documentation
| byte readMap | ( | char * | filename, | |
| void * | data | |||
| ) |
Read data for spatial (map) input binary file.
- Parameters:
-
filename data variable's name data variable's array data
- Returns:
- byte size of data values
Definition at line 43 of file Serial.c.
References debug, dFile, gbl_size, ModelPath, ProjName, and read_header().
Referenced by read_map_file().
00044 { 00045 int gsize, nbytes; byte bsize; 00046 FILE* bFile; 00047 char binFileName[300]; 00048 00049 bsize = read_header(filename); 00050 00051 sprintf(binFileName,"%s/%s/Data/Map_bin/%s",ModelPath,ProjName,filename); 00052 00053 if((bFile = fopen(binFileName,"rb") ) == NULL ) { 00054 fprintf(stderr,"Can't Open Mapfile: %s",binFileName); 00055 exit(0); 00056 } 00057 gsize = gbl_size[0] * gbl_size[1]; 00058 nbytes = fread((char*)data, (int)bsize, gsize, bFile ); 00059 if(debug) { 00060 fprintf(dFile," %d of %d items Read for %s, size = %x bytes ", 00061 nbytes,gsize,filename,bsize); 00062 } 00063 fclose(bFile); 00064 return bsize; 00065 }

| int read_header | ( | char * | filename | ) |
Read header info for spatial (map) input binary file.
- Parameters:
-
filename data variable's file name
- Returns:
- byte size of data values
Definition at line 72 of file Serial.c.
References conv_easting_NAD27toNAD83, conv_northing_NAD27toNAD83, dFile, gbl_size, ModelPath, ProjName, scan_forward(), UTMeast, UTMeastNAD83, UTMnorth, UTMnorthNAD83, UTMsouth, UTMsouthNAD83, UTMwest, and UTMwestNAD83.
Referenced by readMap().
00073 { 00074 FILE* file; 00075 char headerFileName[300]; 00076 00077 int rows, cols, i, Dsize=1, test, offset; 00078 char ch, dirCh; 00079 long int start; 00080 00081 sprintf(headerFileName,"%s/%s/Data/Map_head/%s",ModelPath,ProjName,filename); 00082 00083 file = fopen(headerFileName,"r"); 00084 if(file==NULL) { 00085 fprintf(stderr,"Unable to open header file %s.\n",headerFileName); 00086 fflush(stderr); 00087 exit(0); 00088 } 00089 else { 00090 fprintf(dFile,"Successfully opened header file %s.\n",headerFileName); 00091 fflush(dFile); 00092 } 00093 /* v2.8.2 - get the UTM coordinates of model, for use in self-documenting NetCDF map output */ 00094 scan_forward(file,"north:"); 00095 fscanf(file,"%f",&UTMnorth); 00096 fprintf(dFile,"UTMnorth = %f\n",UTMnorth); 00097 scan_forward(file,"south:"); 00098 fscanf(file,"%f",&UTMsouth); 00099 fprintf(dFile,"UTMsouth = %f\n",UTMsouth); 00100 scan_forward(file,"east:"); 00101 fscanf(file,"%f",&UTMeast); 00102 fprintf(dFile,"UTMeast = %f\n",UTMeast); 00103 scan_forward(file,"west:"); 00104 fscanf(file,"%f",&UTMwest); 00105 fprintf(dFile,"UTMwest = %f\n",UTMwest); 00106 00107 scan_forward(file,"cols:"); 00108 fscanf(file,"%d",&cols); 00109 fprintf(dFile,"cols = %d\n",cols); 00110 scan_forward(file,"rows:"); 00111 fscanf(file,"%d",&rows); 00112 fprintf(dFile,"rows = %d\n",rows); 00113 fflush(dFile); 00114 if( !( rows == gbl_size[0] && cols == gbl_size[1] ) ) { 00115 fprintf(stderr,"Error, Wrong map size: %s: (%d,%d) : (%d,%d)\n", 00116 headerFileName,rows,gbl_size[0],cols,gbl_size[1]); 00117 } 00118 start = ftell(file); 00119 test = scan_forward(file,"format:"); 00120 if(test < 1) { 00121 clearerr(file); 00122 fseek(file, start, 0); 00123 } 00124 else { 00125 fscanf(file,"%d",&Dsize); 00126 fprintf(dFile,"size = %d\n",Dsize); 00127 Dsize = Dsize + 1; /* using header generated by GRASS, the byte size is actually the value read from header plus 1 */ 00128 if( Dsize < 1 || Dsize > 4 ) { 00129 fprintf(stderr,"Error, Illegal map size: %s (%d)\n",headerFileName,Dsize); 00130 exit(0); 00131 } 00132 } 00133 00134 /* v2.8.2 - convert the ELM UTM coordinates from NAD27 to NAD83, for use in the netCDF output */ 00135 UTMnorthNAD83 = UTMnorth + conv_northing_NAD27toNAD83; 00136 UTMsouthNAD83 = UTMsouth + conv_northing_NAD27toNAD83; 00137 UTMeastNAD83 = UTMeast + conv_easting_NAD27toNAD83; 00138 UTMwestNAD83 = UTMwest + conv_easting_NAD27toNAD83; 00139 00140 fclose(file); 00141 return Dsize; 00142 }

| void writeMap | ( | char * | filename, | |
| void * | data, | |||
| int | bSize, | |||
| unsigned char | type, | |||
| int | index | |||
| ) |
Write data for spatial (map) output binary file.
- Parameters:
-
filename Filename (variable name) of data data Data array bSize Byte size of data type Type, or format, of (binary) data index Index used in HDF output
Definition at line 151 of file Serial.c.
References gbl_size, s0, s1, and write_header().
Referenced by write_map_file().
00152 { 00153 int gsize, istat; 00154 FILE* bFile; 00155 00156 if (index == 0 && type=='M') write_header(filename,bSize); 00157 00158 if(type == 'H') { 00159 #if HDF4 00160 if(index==0) { istat = DFR8putimage(filename,(char*)data,s1,s0,0); } 00161 else { istat = DFR8addimage(filename,(char*)data,s1,s0,0); } 00162 if(istat != 0) printf("\nerror writing HDF file: %s\n",filename); 00163 #endif 00164 } 00165 00166 else if(type == 'C') { 00167 printf("\nError in configuring netCDF output, which can only be 4-byte floating point values: %s\n",filename); 00168 exit(0); 00169 } 00170 00171 else { 00172 if((bFile = fopen(filename,"wb") ) == NULL ) { 00173 fprintf(stderr,"Can't Open Mapfile: %s",filename); 00174 exit(0); 00175 } 00176 gsize = gbl_size[0]*gbl_size[1]*bSize; 00177 fwrite((char*)data, gsize, 1, bFile ); 00178 00179 fclose(bFile); 00180 } 00181 }

| void writeFloatMap | ( | char * | filename, | |
| char * | VARNAME, | |||
| float * | data, | |||
| int | bSize, | |||
| unsigned char | type, | |||
| int | index, | |||
| char * | thisvarUnits, | |||
| int | outstep | |||
| ) |
Write floating point data for spatial (map) output binary file (new v2.8.2).
- Parameters:
-
filename Filename (path and variable names) of data VARNAME The variable name of data data Data array of the variable bSize Byte size of data type Type, or format, of (binary) data index Index used in output sequence thisvarUnits Units of the output variable
- Remarks:
- New for v.2.8.2, providing capability to write (4 byte) floating point output maps. Some of this may not be elegant, but will get it done for now. One issue is that the size of the output array is actually s0+2, s1+2, which results in having larger array sizes for these output maps.
Definition at line 197 of file Serial.c.
References gbl_size, write_header(), and writeFloatNetCDF().
Referenced by write_map_file().
00198 { 00199 int gsize; 00200 FILE* bFile; 00201 00202 00203 if(type == 'H') { /* HDF */ 00204 printf("\nError in configuring HDF output, which can only be 1-byte values (you requested 4): %s\n",filename); 00205 exit(0); 00206 } 00207 00208 else if(type == 'C') { /* netCDF */ 00209 writeFloatNetCDF(filename,VARNAME,data,bSize,type,index,thisvarUnits, outstep); 00210 00211 } 00212 else if(type == 'M') { /* generic binary Map */ 00213 if (index == 0) write_header(filename,bSize); 00214 00215 if((bFile = fopen(filename,"wb") ) == NULL ) { 00216 fprintf(stderr,"Can't Open output Mapfile: %s",filename); 00217 exit(0); 00218 } 00219 gsize = (gbl_size[0]+2)*(gbl_size[1]+2)*bSize; /* note size is 2 rows and 2 cols larger than standard input/output arrays */ 00220 fwrite(data, gsize, 1, bFile ); 00221 00222 fclose(bFile); 00223 } 00224 00225 else { /* this error shouldn't be reached, but it is possible */ 00226 printf("\nError in configuring 4-byte output request, which can only be netCDF ('C' argument in 'G' command) or generic map binary ('M' argument in 'G' command): output variable= %s\n",filename); 00227 } 00228 00229 }
| void writeFloatNetCDF | ( | char * | filename, | |
| char * | VARNAME, | |||
| float * | data, | |||
| int | bSize, | |||
| unsigned char | type, | |||
| int | index, | |||
| char * | thisvarUnits, | |||
| int | outstep | |||
| ) |
Definition at line 252 of file Serial.c.
References celWid, simTime::da, ERR, initDateRead, simTime::mo, modelName, modelVers, nc_dataRowRev, NDIMS, offMap_float, s0, s1, SimAlt, SimModif, SimTime, T, simTime::TIME, UTMnorthNAD83, UTMwestNAD83, and simTime::yr.
Referenced by writeFloatMap().
00253 { 00254 int retval; 00255 int ncid, dimids[NDIMS]; 00256 size_t start[NDIMS], count[NDIMS]; 00257 int time_dimid, x_dimid, y_dimid; 00258 int x_id, y_id, time_id, varid, date_id, TM_id; 00259 char s_mo[3], s_da[3], ELMdate[11], dateUnits[100], attribute_string[1000]; 00260 float north[s0+2],east[s1+2]; 00261 float northrev[s0+2]; /* for the reversal of the row ordering in having output w/ origin at lower left */ 00262 00263 time_t begin_sim, end_sim; /* Feb 2011 - just threw in here - poor practice!! */ 00264 struct tm *local_begin_time; /* struct holding real-time calendar time from standard time.h */ 00265 /* a couple of items that use the standard time.h to clock the simulation runtime*/ 00266 begin_sim = time(NULL); 00267 local_begin_time = localtime(&begin_sim); 00268 00269 00270 int i,j, irev; 00271 00272 /* this prevents code from being compiled when netCDF is not installed 00273 (If you do not have netCDF installed on your system, you must set NCDF3 to false in globals.h) */ 00274 #if NCDF3 00275 /* append the date (yyyymmdd) to the root filename*/ 00276 if (SimTime.mo[0]<10) sprintf(s_mo,"0%d", SimTime.mo[0]); 00277 else sprintf(s_mo,"%d", SimTime.mo[0]); 00278 if (SimTime.da[0]<10) sprintf(s_da,"0%d", SimTime.da[0]); 00279 else sprintf(s_da,"%d", SimTime.da[0]); 00280 sprintf(ELMdate,"%d-%s-%s",SimTime.yr[0],s_mo,s_da); 00281 00282 /* Coordinates used for the netCDF cell attributes are the centroids: 00283 float maps are 1-cell larger on each side, so 00284 subtract 1 cell width to start at western edge, and 00285 add 1 cell width to start at northern edge 00286 (remember that origin of SME/ELM is upper left, 00287 while UTM goes in positive direction from lower left) 00288 We then use the centroid (not upper and left edge) of each cell for coords, 00289 so (- celWid + 0.5*celWid) = (-0.5*celWid) */ 00290 for (i=0; i< (s0+2); i++) { 00291 for (j=0; j< (s1+2); j++) { 00292 // east[j] = UTMwest - 0.5 * celWid + (j * celWid); 00293 // north[i]= UTMnorth + 0.5 * celWid - (i * celWid); 00294 // Note that we've switched to having output in NAD83 horizontal datum - if you change this, be sure to alter the metadata strings below 00295 east[j] = UTMwestNAD83 - 0.5 * celWid + (j * celWid); 00296 north[i]= UTMnorthNAD83 + 0.5 * celWid - (i * celWid); 00297 } 00298 } 00299 00300 /* reverse row order (ELM origin is upper left, this outputs origin at lower left) */ 00301 for (i=0; i< (s0+2); i++) { 00302 for (j=0; j< (s1+2); j++) { 00303 irev = (s0+1) - i; 00304 nc_dataRowRev[T(irev,j)] = data[T(i,j)]; 00305 northrev[irev] = north[i]; 00306 } 00307 } 00308 00309 /* Create the file. The NC_CLOBBER parameter tells netCDF to 00310 * overwrite this file, if it already exists; NC_NOCLOBBER does not overwrite. */ 00311 if (index==0) { 00312 if ( (retval = nc_create(filename, NC_CLOBBER, &ncid))) 00313 ERR(retval); 00314 00315 /* Define the dimensions: x, y, and time outstep-number (sequence). NetCDF will hand back an ID for each. */ 00316 if ((retval = nc_def_dim(ncid, "time", NC_UNLIMITED, &time_dimid))) 00317 ERR(retval); 00318 if ((retval = nc_def_dim(ncid, "y", (s0+2), &y_dimid))) /* note the s0+2 array size */ 00319 ERR(retval); 00320 if ((retval = nc_def_dim(ncid, "x", (s1+2), &x_dimid))) /* note the s1+2 array size */ 00321 ERR(retval); 00322 00323 /* The dimids array is used to pass the IDs of the dimensions of 00324 * the variable. */ 00325 dimids[0] = time_dimid; 00326 dimids[1] = y_dimid; 00327 dimids[2] = x_dimid; 00328 00329 /* Define the model output variable, time, x, y, and transverse_mercator (required for EverVIEW application) variables. 00330 The type of the model output variable in this case is * NC_FLOAT (4-byte floating point). */ 00331 if ((retval = nc_def_var(ncid, "transverse_mercator", NC_CHAR, 0, 00332 0, &TM_id))) 00333 { printf("oops: %s",nc_strerror(retval) ); exit(-1); } 00334 00335 if ((retval = nc_def_var(ncid, VARNAME, NC_FLOAT, NDIMS, 00336 dimids, &varid))) 00337 ERR(retval); 00338 if ((retval = nc_def_var(ncid, "time", NC_FLOAT, 1, 00339 &time_dimid, &time_id))) 00340 ERR(retval); 00341 if ((retval = nc_def_var(ncid, "y", NC_FLOAT, 1, 00342 &y_dimid, &y_id))) 00343 ERR(retval); 00344 if ((retval = nc_def_var(ncid, "x", NC_FLOAT, 1, 00345 &x_dimid, &x_id))) 00346 ERR(retval); 00347 00348 /* #### Attributes of the time variable. #### */ 00349 if ((retval = nc_put_att_text(ncid, time_id, "long_name", 00350 strlen("model time"), "model time"))) 00351 ERR(retval); 00352 /* Assign units attributes to the julian time. In definition mode, the current date is initial day 0 of simulation */ 00353 sprintf(dateUnits,"days since %sT12:00:00Z",ELMdate); 00354 if ((retval = nc_put_att_text(ncid, time_id, "units", 00355 strlen(dateUnits), dateUnits))) 00356 ERR(retval); 00357 if ((retval = nc_put_att_text(ncid, time_id, "_CoordinateAxisType", 00358 strlen("Time"), "Time"))) 00359 ERR(retval); 00360 00361 00362 /* #### Attributes of the model output variable. #### */ 00363 /* Assign "long_name" attribute to the variable - to do this right, need to pass the variable's description into here. */ 00364 if ((retval = nc_put_att_text(ncid, varid, "long_name", 00365 strlen(VARNAME), VARNAME))) 00366 ERR(retval); 00367 if ((retval = nc_put_att_text(ncid, varid, "units", 00368 strlen(thisvarUnits), thisvarUnits))) 00369 ERR(retval); 00370 if ((retval = nc_put_att_float(ncid, varid, "missing_value", /* note that "_FillValue" is updated attribute, but EverVIEW didn't respond to that, so using deprecated "missing_value" */ 00371 NC_FLOAT, 1, &offMap_float))) 00372 ERR(retval); 00373 if ((retval = nc_put_att_int(ncid, varid, "OutputInterval_inDays", 00374 NC_INT, 1, &outstep))) 00375 ERR(retval); 00376 if ((retval = nc_put_att_text(ncid, varid, "coordinates", 00377 strlen("x y"), "x y"))) 00378 ERR(retval); 00379 if ((retval = nc_put_att_text(ncid, varid, "grid_mapping", 00380 strlen("transverse_mercator"), "transverse_mercator"))) 00381 ERR(retval); 00382 00383 /* for compatability with other south Florida models, we switched (only the netCDF) output to NAD83 (in the above spatial loop) */ 00384 sprintf(attribute_string,"PROJCS[\"NAD_1983_UTM_Zone_17N\",GEOGCS[\"GCS_North_American_1983\",DATUM[\"D_North_American_1983\",SPHEROID[\"GRS_1980\",6378137.0,298.257222101]],PRIMEM[\"Greenwich\",0.0],UNIT[\"Degree\",0.0174532925199433]],PROJECTION[\"Transverse_Mercator\"],PARAMETER[\"False_Easting\",500000.0],PARAMETER[\"False_Northing\",0.0],PARAMETER[\"Central_Meridian\",-81.0],PARAMETER[\"Scale_Factor\",0.9996],PARAMETER[\"Latitude_Of_Origin\",0.0],UNIT[\"Meter\",1.0]]"); 00385 00386 if ((retval = nc_put_att_text(ncid, varid, "esri_pe_string", 00387 strlen(attribute_string), attribute_string))) 00388 ERR(retval); 00389 00390 /* #### Attributes of the UTM coordinate variables. #### */ 00391 if ((retval = nc_put_att_text(ncid, x_id, "long_name", 00392 strlen("easting (x) coordinate"), "easting (x) coordinate"))) 00393 ERR(retval); 00394 if ((retval = nc_put_att_text(ncid, y_id, "long_name", 00395 strlen("northing (y) coordinate"), "northing (y) coordinate"))) 00396 ERR(retval); 00397 if ((retval = nc_put_att_text(ncid, x_id, "standard_name", 00398 strlen("projection_x_coordinate"), "projection_x_coordinate"))) 00399 ERR(retval); 00400 if ((retval = nc_put_att_text(ncid, y_id, "standard_name", 00401 strlen("projection_y_coordinate"), "projection_y_coordinate"))) 00402 ERR(retval); 00403 if ((retval = nc_put_att_text(ncid, x_id, "units", 00404 strlen("m"), "m"))) 00405 ERR(retval); 00406 if ((retval = nc_put_att_text(ncid, y_id, "units", 00407 strlen("m"), "m"))) 00408 ERR(retval); 00409 if ((retval = nc_put_att_text(ncid, x_id, "_CoordinateAxisType", 00410 strlen("GeoX"), "GeoX"))) 00411 ERR(retval); 00412 if ((retval = nc_put_att_text(ncid, y_id, "_CoordinateAxisType", 00413 strlen("GeoY"), "GeoY"))) 00414 ERR(retval); 00415 if ((retval = nc_put_att_text(ncid, x_id, "CellRegistration", 00416 strlen("coords are at cell centroids"), "coords are at cell centroids"))) 00417 ERR(retval); 00418 if ((retval = nc_put_att_text(ncid, y_id, "CellRegistration", 00419 strlen("coords are at cell centroids"), "coords are at cell centroids"))) 00420 ERR(retval); 00421 00422 /* #### Attributes of the transverse_mercator variable (done only for EverVIEW application). #### */ 00423 if ((retval = nc_put_att_text(ncid, TM_id, "grid_mapping_name", 00424 strlen("transverse_mercator"), "transverse_mercator"))) 00425 ERR(retval); 00426 if ((retval = nc_put_att_text(ncid, TM_id, "longitude_of_central_meridian", 00427 strlen(" -81."), " -81."))) 00428 ERR(retval); 00429 if ((retval = nc_put_att_text(ncid, TM_id, "latitude_of_projection_origin", 00430 strlen("0."), "0."))) 00431 ERR(retval); 00432 if ((retval = nc_put_att_text(ncid, TM_id, "scale_factor_at_central_meridian", 00433 strlen("0.9996"), "0.9996"))) 00434 ERR(retval); 00435 if ((retval = nc_put_att_text(ncid, TM_id, "false_easting", 00436 strlen("500000."), "500000."))) 00437 ERR(retval); 00438 if ((retval = nc_put_att_text(ncid, TM_id, "false_northing", 00439 strlen("0."), "0."))) 00440 ERR(retval); 00441 if ((retval = nc_put_att_text(ncid, TM_id, "_CoordinateTransformType", 00442 strlen("Projection"), "Projection"))) 00443 ERR(retval); 00444 if ((retval = nc_put_att_text(ncid, TM_id, "_CoordinateAxisTypes", 00445 strlen("GeoX GeoY"), "GeoX GeoY"))) 00446 ERR(retval); 00447 if ((retval = nc_put_att_text(ncid, TM_id, "units", 00448 strlen("m"), "m"))) 00449 ERR(retval); 00450 00451 00452 /* #### Global attributes. #### */ 00453 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "title", 00454 strlen("Everglades Landscape Model output"), "Everglades Landscape Model output"))) 00455 ERR(retval); 00456 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "ModelName", 00457 strlen(modelName), modelName))) 00458 ERR(retval); 00459 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "ModelVersion", 00460 strlen(modelVers), modelVers))) 00461 ERR(retval); 00462 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "ScenarioType", 00463 strlen(SimAlt), SimAlt))) 00464 ERR(retval); 00465 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "Scenario", 00466 strlen(SimModif), SimModif))) 00467 ERR(retval); 00468 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "Day0_SimulationPeriod", 00469 strlen(initDateRead), initDateRead))) 00470 ERR(retval); 00471 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "Time_model_run_started", 00472 strlen(asctime(local_begin_time)), asctime(local_begin_time)))) 00473 ERR(retval); 00474 /* todo get rid of this hardcoding */ 00475 sprintf(attribute_string,"H. Carl Fitz, carlfitz3@gmail.com, EcoLandMod, Inc, http://www.ecolandmod.com"); 00476 00477 if ((retval = nc_put_att_text(ncid, NC_GLOBAL, "modelDeveloper", 00478 strlen(attribute_string), attribute_string))) 00479 ERR(retval); 00480 00481 00482 /* #### End define mode. This tells netCDF we are done defining metadata. #### */ 00483 if ((retval = nc_enddef(ncid))) 00484 ERR(retval); 00485 00486 00487 00488 /* Still on the first outstep, write the coordinate variable data. This will put the northings 00489 and eastings of our data grid into the netCDF file. */ 00490 if ((retval = nc_put_var_float(ncid, y_id, &northrev[0]))) /* note the use of reverse rows */ 00491 ERR(retval); 00492 if ((retval = nc_put_var_float(ncid, x_id, &east[0]))) 00493 ERR(retval); 00494 00495 if ((retval = nc_close(ncid))) 00496 ERR(retval); 00497 00498 00499 } /* end of all defs. Done once, prior to first writing of data; have a closed file */ 00500 00501 /* These settings tell netcdf to write one timestep of data. (The 00502 value of start[0] below tells netCDF which (index) 00503 timestep to write.) */ 00504 count[0] = 1; 00505 count[1] = s0+2; /* was s0+2 */ 00506 count[2] = s1+2; /* was s1+2 */ 00507 start[1] = 0; 00508 start[2] = 0; 00509 00510 start[0] = index; /* tried to use 'SimTime.TIME' , but it filled in multiple blank days */ 00511 /* now open the ncdf file, and write to it */ 00512 if ((retval = nc_open(filename, NC_WRITE, &ncid))) 00513 ERR(retval); 00514 if ((retval = nc_inq_varid(ncid, VARNAME, &varid))) 00515 ERR(retval); 00516 if ((retval = nc_inq_varid(ncid, "time", &time_id))) 00517 ERR(retval); 00518 00519 /* Write the data to the file. */ 00520 if ((retval = nc_put_vara_float(ncid, varid, start, count, nc_dataRowRev))) 00521 ERR(retval); 00522 if ((retval = nc_put_var1_float(ncid, time_id, start, &SimTime.TIME ))) 00523 ERR(retval); 00524 00525 /* Close the file. This frees up any internal netCDF resources 00526 * associated with the file, and flushes any buffers. */ 00527 if ((retval = nc_close(ncid))) 00528 ERR(retval); 00529 00530 /* end of netCDF code */ 00531 #else 00532 00533 printf("\nError in configuring 4-byte output request, you asked for netCDF ('C' argument in 'G' command) output, but your globals.h file has NCDF3 set to false: output variable= %s\n",filename); 00534 exit(0); 00535 00536 #endif 00537 }
| void write_header | ( | char * | mapFileName, | |
| int | size | |||
| ) |
Write header info for spatial (map) output binary file.
- Parameters:
-
mapFileName Filename (variable name) of data size Byte size of data
Definition at line 545 of file Serial.c.
References celWid, dFile, gbl_size, UTMeast, UTMnorth, UTMsouth, and UTMwest.
Referenced by writeFloatMap(), and writeMap().
00546 { 00547 FILE* header; 00548 char headFIleName[320]; 00549 int outrow,outcol; 00550 /* v2.8.2 modified - GRASS format header */ 00551 sprintf(headFIleName,"%s_header.txt",mapFileName); 00552 fprintf(dFile,"Writing Header: filename = %s\n",headFIleName); 00553 fflush(dFile); 00554 header = fopen(headFIleName,"w"); 00555 if(header==NULL) { 00556 fprintf(stderr,"Can't open header file: %s\n",headFIleName); 00557 fflush(stderr); exit(-1); 00558 } 00559 if(size == 4) { /* v2.8.2 - size of float arrays are 2 larger (both row and col) than the standard output */ 00560 outrow = gbl_size[0]+2; 00561 outcol = gbl_size[1]+2; 00562 } 00563 else { 00564 outrow = gbl_size[0]; 00565 outcol = gbl_size[1]; 00566 } 00567 /* GRASS header - note that the "format" byte size is the size - 1 . Also, the UTM zone is hardcoded to 17 here - lazy (unused anywhere in model) */ 00568 fprintf(header,"proj: 1\nzone: 17\nnorth: %.0f\nsouth: %.0f\neast: %.0f\nwest: %.0f\n", 00569 UTMnorth, UTMsouth, UTMeast, UTMwest); 00570 fprintf(header,"cols: %d\nrows: %d\ne-w resol: %.0f\nn-s resol: %.0f\nformat: %d\ncompressed: 0\n", 00571 outcol,outrow,celWid,celWid,(size-1) ); 00572 fclose(header); 00573 }
| float* readSeriesCol | ( | char * | filename, | |
| int | format, | |||
| int | index, | |||
| int * | Npt, | |||
| float * | TStep, | |||
| int | col | |||
| ) |
(Unused). Read time series input data for spatial interpolations
- Parameters:
-
filename Filename of data format Always float data format index Unused. Was used to concatenate multiple files over time Npt Number of time points TStep Time step col column ID (for different variables)
Definition at line 585 of file Serial.c.
References debug, dFile, Jdate_init, julday(), nalloc(), PORnumday, and scan_forward().
Referenced by PTS_SetFields().
00586 { 00587 int line = 0, cread=1, itest, j, sLen = 372 ; 00588 FILE *cfile; 00589 char ctest, cmark[373], ret = '\n'; 00590 unsigned char cmove[373], cmv=0; 00591 char tfilename[200], date_read[20],ss[200]; 00592 int yyyy,mm, dd; 00593 double Jdate_read; 00594 float *tvar, *nullvar, first[10]; 00595 00596 sprintf(tfilename,"%s%d.ts",filename,index); 00597 if(debug>4) fprintf(dFile,"\nReading file %s, col %d, index = %d\n",tfilename,col,index); fflush(dFile); 00598 cfile = fopen(tfilename,"r"); 00599 if(cfile==NULL) { 00600 if( index > 0 ) { fprintf(stdout,"\nWARNING: Unable to open timeseries file %s, using %s0.ts\n",tfilename,filename); return 0; } 00601 else { fprintf(stdout,"\nERROR: Unable to open timeseries file %s\n",tfilename); exit(-1); } 00602 } 00603 00604 if (format != 2) { fprintf(stderr,"ERROR: only using floats in read_timeseries: %d\n",format); exit(0); } 00605 00606 sLen = PORnumday; /* read/determined in genericDriver */ 00607 tvar = (float*) nalloc( (sLen+2)*sizeof(float), "timeseries temp" ); 00608 scan_forward(cfile,"{END COMMENT}\n"); 00609 fgets(ss,200,cfile); /* skip the header line */ 00610 00611 do { 00612 fscanf(cfile, "%d %d %d %f %f %f %f", &yyyy,&mm,&dd,&first[1],&first[2],&first[3],&first[4] ); 00613 /* julian day, returns a double (args = month, day, year, hours, minutes, seconds) */ 00614 Jdate_read = julday(mm, dd, yyyy, 0, 0, 0); 00615 if (Jdate_read > Jdate_init) {printf (" \n***Error\nInit date in %s file > simulation start date.\n",tfilename); exit (-1);} 00616 if (feof(cfile)) { 00617 printf (" \n***Error\nNon-matching starting date for %s meteorological time series\n",tfilename); 00618 exit (-1); 00619 } 00620 } while (Jdate_read < Jdate_init); 00621 00622 tvar[0] = first[col]; /* have now read the first data record, assign to array */ 00623 if(debug>4) fprintf(dFile,"%d %d %d %f\n",yyyy,mm,dd,tvar[0]); 00624 for (line = 1; line<sLen; line++) { 00625 00626 /* each case for reading a different column */ 00627 switch (col) { 00628 case 1: 00629 itest = fscanf(cfile,"%d %d %d %f %f %f %f",&yyyy,&mm,&dd, &(tvar[ line ]),&nullvar, &nullvar, &nullvar); 00630 break; 00631 case 2: 00632 itest = fscanf(cfile,"%d %d %d %f %f %f %f",&yyyy,&mm,&dd,&nullvar, &(tvar[ line ]), &nullvar, &nullvar); 00633 break; 00634 case 3: 00635 itest = fscanf(cfile,"%d %d %d %f %f %f %f",&yyyy,&mm,&dd,&nullvar, &nullvar, &(tvar[ line ]), &nullvar); 00636 break; 00637 case 4: 00638 itest = fscanf(cfile,"%d %d %d %f %f %f %f",&yyyy,&mm,&dd,&nullvar, &nullvar, &nullvar, &(tvar[ line ])); 00639 break; 00640 default: 00641 printf ("\nError in interpolation time series data.\n"); 00642 exit(-1); 00643 break; 00644 } /* end switch */ 00645 00646 if(debug>4) fprintf(dFile,"%d %d %d %f\n",yyyy,mm,dd,tvar[ line ]); 00647 if (feof(cfile)) {printf (" \n***Error\nExceeded number of records for %s meteorological time series\n",tfilename); exit (-1); } 00648 } 00649 tvar[ sLen ] = *TStep; 00650 00651 if(debug>4) { fprintf(dFile,"\nDone Reading file %s\n",tfilename); fflush(dFile); } 00652 00653 fflush(stdout); 00654 fclose(cfile); 00655 *Npt = sLen; 00656 return tvar; 00657 }
| float* get_hab_parm | ( | char * | s_parm_name, | |
| int | s_parm_relval, | |||
| char * | parmName | |||
| ) |
Read the input data file to get a habitat-specfic parameter array.
This gets an array of the targeted parameter, a value for each of (usually) multiple habitats.
- Parameters:
-
s_parm_name Name of the sensitivity parameter being varied for this run s_parm_relval Indicator of current value of relative range in sensitivity parm values: nominal (0), low (1), or high (2) parmName String that identifies the parameter to be read in dbase file
- Returns:
- fTmp Array of values of single parameter across multiple habitats
Definition at line 671 of file Serial.c.
References dFile, EOL, EOS, False, get_Nth_parm(), gHabChar, goto_index(), habNumTot, match_Sparm(), MAX_NHAB, modelFileName, ModelPath, msgStr, ProgExec, ProjName, prog_attr::S_ParmVal, Scip(), TAB, True, usrErr(), and WriteMsg().
Referenced by ReadHabParms().
00672 { 00673 int hIndex, end=0; 00674 FILE* parmFile; 00675 float* fTmp; 00676 char lline[2001], *line; 00677 int pLoc = -1; 00678 int foundParm= False; 00679 char parmNameHead[30]; 00680 char modelFileName[300]; 00681 char HabParmFile[300]; 00682 char *fileAppend; 00683 extern ProgAttr *ProgExec; 00684 00685 00686 if ( (fTmp = (float*) malloc( MAX_NHAB * sizeof(float) ) ) == NULL) { 00687 sprintf(msgStr, "Failed to allocate memory for HabParm array.\n ") ; 00688 usrErr(msgStr); 00689 exit( -2 ) ; 00690 }; 00691 fTmp[0] = 0.0; 00692 00693 fileAppend = match_Sparm( s_parm_name, s_parm_relval, parmName); 00694 sprintf(HabParmFile,"HabParms_%s",fileAppend); 00695 sprintf(modelFileName,"%s/%s/Data/%s",ModelPath,ProjName,HabParmFile); 00696 parmFile = fopen(modelFileName,"r"); 00697 if(parmFile==NULL) {fprintf(stdout,"\nERROR: Unable to open dBase file %s\n",modelFileName); exit(-1); } 00698 00699 fgets(lline, 2000, parmFile); /* skip 1st header line */ 00700 fgets(lline, 2000, parmFile); /* read 2nd header line with column names */ 00701 line=lline; 00702 00703 while (!foundParm) 00704 { 00705 sscanf( line,"%s",&parmNameHead); 00706 if (strcmp(parmName,parmNameHead) ==0) foundParm = True; 00707 pLoc++; 00708 if (*line == EOS || *line == EOL) { 00709 sprintf(msgStr,"ERROR: Could not find header tag of %s in HabParms dbase file.", parmName); 00710 usrErr(msgStr); 00711 exit(-1); 00712 } 00713 line = Scip( line, TAB ); 00714 } 00715 00716 sprintf(msgStr,"Parameter group: %s",parmName); 00717 fflush(dFile); 00718 WriteMsg(msgStr,1); 00719 for( hIndex = 1; hIndex < MAX_NHAB; hIndex++) { 00720 goto_index (parmFile, gHabChar, hIndex); 00721 fTmp[hIndex] = get_Nth_parm( parmFile, pLoc, &end, hIndex ); 00722 if(end) break; 00723 } 00724 fclose(parmFile); 00725 habNumTot = hIndex-1; 00726 00727 fflush(dFile); 00728 /* TODO: this HARDCODES into the BIR (BIRavg) model output file ONLY, 00729 the incorrect impression of running sensitivity on a single habitat ID 00730 (hab 2 = sawgrass plain, ELMv2.4) */ 00731 if (strcmp(s_parm_name,parmName) == 0) ProgExec->S_ParmVal = fTmp[2]; 00732 return fTmp; 00733 }
| char* match_Sparm | ( | char * | s_parm_name, | |
| int | s_parm_relval, | |||
| char * | parmName | |||
| ) |
Determine if the parameter is being varied for a sensitivity analysis.
- Parameters:
-
s_parm_name Name of the sensitivity parameter being varied for this run s_parm_relval Indicator of current value of relative range in sensitivity parm values: nominal (0), low (1), or high (2) parmName String that identifies the parameter to be read in dbase file
- Returns:
- fileAppend The suffix (not extension) to be appended to end of base name of parameter input file.
Definition at line 742 of file Serial.c.
References msgStr, and usrErr().
Referenced by get_global_parm(), get_hab_parm(), and get_modexperim_parm().
00743 { 00744 char *fileAppend; 00745 00746 if (strcmp(s_parm_name,parmName) == 0) /* match, it is a parameter being varied in this set of runs */ 00747 { 00748 switch (s_parm_relval) { 00749 case 0: fileAppend = "NOM"; break; 00750 case 1: fileAppend = "LO"; break; 00751 case 2: fileAppend = "HI"; break; 00752 default: { 00753 sprintf(msgStr,"ERROR: The relative parameter value (%d) is unknown for %s",s_parm_relval, s_parm_name); 00754 usrErr(msgStr); 00755 exit(-1); 00756 } 00757 } 00758 } 00759 else 00760 { /* no match, it is NOT a parameter being varied in this set of runs */ 00761 fileAppend = "NOM"; 00762 } 00763 return (fileAppend); 00764 }

| float get_global_parm | ( | char * | s_parm_name, | |
| int | s_parm_relval, | |||
| char * | parmName | |||
| ) |
Read the input data file to get a global parameter.
This gets the single global value of the targeted parameter.
- Parameters:
-
s_parm_name Name of the sensitivity parameter being varied for this run s_parm_relval Indicator of current value of relative range in sensitivity parm values: nominal (0), low (1), or high (2) parmName String that identifies the parameter to be read in dbase file
- Returns:
- parmVal The value of the global parameter
Definition at line 775 of file Serial.c.
References Exit(), match_Sparm(), modelFileName, ModelPath, msgStr, ProgExec, ProjName, prog_attr::S_ParmVal, scan_forward(), usrErr(), and WriteMsg().
Referenced by ReadGlobalParms().
00776 { 00777 int test; 00778 char modelFileName[300]; 00779 FILE *parmFile; 00780 float parmVal; 00781 char parmNameHead[30]; 00782 extern ProgAttr *ProgExec; 00783 00784 00785 char GlobalParmFile[50]; 00786 char *fileAppend; 00787 00788 fileAppend = match_Sparm( s_parm_name, s_parm_relval, parmName); 00789 sprintf(GlobalParmFile,"GlobalParms_%s",fileAppend); 00790 sprintf(modelFileName,"%s/%s/Data/%s",ModelPath,ProjName,GlobalParmFile); 00791 00792 parmFile = fopen(modelFileName,"r"); 00793 if(parmFile==NULL) { 00794 sprintf(msgStr,"ERROR, can't open data file %s",modelFileName); 00795 usrErr(msgStr); 00796 Exit(0); 00797 } 00798 00799 sprintf(parmNameHead,"%s=", parmName); 00800 scan_forward(parmFile,parmNameHead); 00801 if ( (test = fscanf(parmFile,"%f",&parmVal) ) <= 0 ) { 00802 sprintf(msgStr,"ERROR in reading %s from %s; see Driver0.out debug file.", parmName, modelFileName); 00803 usrErr(msgStr); 00804 Exit(0); 00805 } 00806 sprintf(msgStr,"%s %f\n",parmNameHead, parmVal); 00807 WriteMsg(msgStr,1); 00808 00809 fclose(parmFile); 00810 if (strcmp(s_parm_name,parmName) == 0) ProgExec->S_ParmVal = parmVal; 00811 return parmVal; 00812 }
| float get_modexperim_parm | ( | char * | s_parm_name, | |
| int | s_parm_relval, | |||
| char * | parmName | |||
| ) |
Read the input data file to get a model experiment parameter.
This gets the single global value of the targeted parameter for special model experiments. (Added for ELM v2.6).
- Parameters:
-
s_parm_name Name of the sensitivity parameter being varied for this run s_parm_relval Indicator of current value of relative range in sensitivity parm values: nominal (0), low (1), or high (2) parmName String that identifies the parameter to be read in dbase file
- Returns:
- parmVal The value of the (global) model experiment parameter
Definition at line 822 of file Serial.c.
References Exit(), match_Sparm(), modelFileName, ModelPath, msgStr, ProgExec, ProjName, prog_attr::S_ParmVal, scan_forward(), usrErr(), and WriteMsg().
Referenced by ReadModExperimParms().
00823 { 00824 int test; 00825 char modelFileName[300]; 00826 FILE *parmFile; 00827 float parmVal; 00828 char parmNameHead[30]; 00829 extern ProgAttr *ProgExec; 00830 00831 00832 char ModExParmFile[50]; 00833 char *fileAppend; 00834 00835 fileAppend = match_Sparm( s_parm_name, s_parm_relval, parmName); 00836 sprintf(ModExParmFile,"ModExperimParms_%s",fileAppend); 00837 sprintf(modelFileName,"%s/%s/Data/%s",ModelPath,ProjName,ModExParmFile); 00838 00839 parmFile = fopen(modelFileName,"r"); 00840 if(parmFile==NULL) { 00841 sprintf(msgStr,"ERROR, can't open data file %s",modelFileName); 00842 usrErr(msgStr); 00843 Exit(0); 00844 } 00845 00846 sprintf(parmNameHead,"%s=", parmName); 00847 scan_forward(parmFile,parmNameHead); 00848 if ( (test = fscanf(parmFile,"%f",&parmVal) ) <= 0 ) { 00849 sprintf(msgStr,"ERROR in reading %s from %s; see Driver0.out debug file.", parmName, modelFileName); 00850 usrErr(msgStr); 00851 Exit(0); 00852 } 00853 sprintf(msgStr,"%s %f\n",parmNameHead, parmVal); 00854 WriteMsg(msgStr,1); 00855 00856 fclose(parmFile); 00857 if (strcmp(s_parm_name,parmName) == 0) ProgExec->S_ParmVal = parmVal; 00858 return parmVal; 00859 }
| void open_point_lists | ( | SeriesParm * | pSeries, | |
| int | nSeries | |||
| ) |
Point time series output: open files and print headers.
Provides for multi-column (locations), multi-file (model variables)format of point time series. The user can define (via Model.outList commands), for any variable in the model, one or more cell (point) locations for which point time series data are written. There can be multiple point TS each for different variables. Thus, we make a separate file for each variable (unknown until user chooses) that has multiple columns for multiple points (unknown until user chooses).
- Parameters:
-
pSeries An array of SeriesParm structs for point time series output nSeries Number of point time series occurances
- Remarks:
- A structure should/could be set up for this, but to maintain minimal changes between the parallel and serial versions of the code (ELMv1.0), for now we just maintain the original data structure and use some counters below to keep track of diff files and numbers of columns (also allowing different output steps as chosen by the user). (Note: The original SME code printed all variables in one column in one file).
- Note:
- TODO: this whole point-time-series output functionality needs improvement (not high priority, Jan 2005). One of the more important capabilities is to have ability to have irregular, calendar-month/year etc output intervals.
Definition at line 881 of file Serial.c.
References cell_pts, seriesParm::Loc, modelName, modelVers, msgStr, numPtFiles, OutputPath, ProjName, SimAlt, SimModif, point2D::x, and point2D::y.
Referenced by main().
00882 { 00883 FILE *oFile; 00884 int i, j, ix, iy, k= 0 ; 00885 00886 for (i=0; i<nSeries; i++) { 00887 if( pSeries[i].data == NULL ) { 00888 if ( numPtFiles>0 ) fclose(oFile); /* in (rare?) case where user asks for no point output, no files stream to close */ 00889 return; 00890 } 00891 if (strcmp(pSeries[i].name,pSeries[i-1].name) != 0) { 00892 /* this is new variable, thus new file. numPtFiles== increment counter of number of variables (one file per variable) used for output */ 00893 if (i>0) { 00894 numPtFiles++; 00895 fclose (oFile); 00896 } 00897 00898 sprintf(msgStr,"%s%s/Output/PtSer/%s.pts",OutputPath,ProjName,pSeries[i].name); 00899 if( (oFile = fopen(msgStr,"w") ) == NULL) { 00900 fprintf(stderr,"\nERROR, unable to open %s point time series output file.",msgStr); 00901 exit(-1); 00902 } 00903 fprintf (oFile, "%s %s %s %s scenario: \nDate\t", &modelName, &modelVers, &SimAlt, &SimModif); 00904 00905 } 00906 /* populate the grid cell locations in the header for this variable/file */ 00907 ix = pSeries[i].Loc.x; 00908 iy = pSeries[i].Loc.y; 00909 fprintf(oFile,"%s:(%d,%d)\t",pSeries[i].name,ix,iy); 00910 cell_pts[numPtFiles]++; /* count the number of cell point locs per variable (1 var per file) */ 00911 00912 } 00913 }
| void send_point_lists2 | ( | SeriesParm * | pSeries, | |
| int | nSeries | |||
| ) |
Point time series output: print time series data to file(s).
open_point_lists Has documentation relevant to this function
- Parameters:
-
pSeries A struct of the variables to have point time series output nSeries Number of point time series occurances
Definition at line 924 of file Serial.c.
References calcdate(), cell_pts, Jdate_init, seriesParm::laststep, seriesParm::Length, msgStr, numPtFiles, OutputPath, seriesParm::outstep, and ProjName.
Referenced by main().
00925 { 00926 FILE *oFile; 00927 int i=0, ii, j, ix, iy, k= 0, last_pt; 00928 int yr_pt[2],mo_pt[2],da_pt[2],hr_pt[2],mi_pt[2]; 00929 double se_pt[2]; 00930 double Jdate_pt; 00931 00932 if (numPtFiles==0) return; 00933 /* k = file/variable counter; i = indiv cellLoc time series counter */ 00934 for (k = 0; k <= numPtFiles; k++) { /* loop over the number of point ts files */ 00935 sprintf(msgStr,"%s%s/Output/PtSer/%s.pts",OutputPath,ProjName,pSeries[i].name); 00936 if( (oFile = fopen(msgStr,"a") ) == NULL) { 00937 fprintf(stderr,"\nERROR, unable to open %s point time series output file.",msgStr); 00938 exit(-1); 00939 } 00940 00941 for(j=pSeries[i].laststep; j<pSeries[i].Length; j++ ) { /*temporal loop */ 00942 Jdate_pt = Jdate_init+j*pSeries[i].outstep; 00943 calcdate( Jdate_pt, mo_pt, da_pt, yr_pt, hr_pt, mi_pt, se_pt); /* get the calendar date info from the julian date */ 00944 fprintf(oFile,"\n%d/%d/%d\t",yr_pt[0],mo_pt[0],da_pt[0] ); /* calendar date in col 1 */ 00945 00946 last_pt = i+cell_pts[k]; /* # of last cellLoc point of the current variable's file */ 00947 for (ii=i; ii<last_pt; ii++) {/* loop over number of point locations per file */ 00948 fprintf(oFile,"%f\t",pSeries[ii].data[j]); 00949 } 00950 } 00951 fclose (oFile); 00952 pSeries[i].laststep = j; /* remember the last temporal outstep for this file */ 00953 i += cell_pts[k]; /* increment the indiv cellLoc time series counter */ 00954 } 00955 return; 00956 }

| void writeSeries | ( | void * | fValue, | |
| char * | label, | |||
| char * | desc, | |||
| int | N0, | |||
| int | N1, | |||
| byte | Mtype, | |||
| byte | format | |||
| ) |
Write to a spatial data window in a debug file.
- Parameters:
-
fValue Model variable (local) array data label Output message label (w/ variable name) desc Message describing the output N0 Range in x (row!) values N1 Range in y (col!) values Mtype General output format type format Numeric format type
Definition at line 970 of file Serial.c.
References cTemp, Driver_outfile, HDF_VERIFY, OutputPath, and ProjName.
Referenced by writeWindow().
00971 { 00972 /* check on these! (who wrote this? Why? (HCF Dec'04) )*/ 00973 int ix, iy; 00974 static unsigned char first_write = 1; 00975 long int ret, dimsizes[2]; 00976 unsigned short int refnum; 00977 if(format == 'H') { 00978 00979 /* NOTE: June 2008 (v2.8.2): do not #define HDF in globals.h to true until until the code here is updated - 00980 this code needs to be updated to HDF4 before it will compile and be used */ 00981 #if HDF 00982 dimsizes[0] = N0; 00983 dimsizes[1] = N1; 00984 sprintf(cTemp,"%s%s:Output:Windows.hdf",OutputPath,ProjName); 00985 DFSDclear(); 00986 00987 switch(Mtype) { 00988 case 'f': 00989 DFSDsetNT(DFNT_FLOAT32); break; 00990 case 'L': 00991 DFSDsetNT(DFNT_FLOAT32); break; 00992 case 'E': 00993 DFSDsetNT(DFNT_FLOAT32); break; 00994 case 'd': case 'i': 00995 DFSDsetNT(DFNT_INT32); break; 00996 case 'c': 00997 DFSDsetNT(DFNT_UINT8); break; 00998 } 00999 if(first_write) { 01000 ret = DFSDputdata(cTemp,2,dimsizes,fValue); 01001 first_write = 0; 01002 } else { 01003 ret = DFSDadddata(cTemp,2,dimsizes,fValue); 01004 } 01005 HDF_VERIFY("DFSDadddata"); 01006 refnum = DFSDlastref(); 01007 ret = DFANputlabel(cTemp,DFTAG_SDG,refnum,label); 01008 HDF_VERIFY("DFANputlabel"); 01009 ret = DFANputdesc(cTemp,DFTAG_SDG,refnum,desc,strlen(desc)); 01010 HDF_VERIFY("DFANputdesc"); 01011 #endif 01012 } 01013 else { 01014 fprintf(Driver_outfile,"\n_________%s\n",label); 01015 fprintf(Driver_outfile,"%s\n",desc); 01016 for (ix=0; ix<N0; ix++) { 01017 fprintf(Driver_outfile,"\n"); 01018 for (iy=0; iy<N1; iy++) { 01019 switch(Mtype) { 01020 case 'f': 01021 if(format=='L') fprintf(Driver_outfile,"%f ",*(((float*)fValue)+iy+ix*N1)); 01022 else if(format=='E') fprintf(Driver_outfile,"%.3E ",*(((float*)fValue)+iy+ix*N1)); 01023 else fprintf(Driver_outfile,"%.3f ",*(((float*)fValue)+iy+ix*N1)); 01024 break; 01025 case 'd': case 'i': 01026 fprintf(Driver_outfile,"%d ",*(((int*)fValue)+iy+ix*N1)); 01027 break; 01028 case 'c': 01029 fprintf(Driver_outfile,"%x ",*(((UCHAR*)fValue)+iy+ix*N1)); 01030 break; 01031 } 01032 } 01033 } 01034 } 01035 }
| void Combine | ( | float * | fValue, | |
| char * | label, | |||
| int | nComp, | |||
| int | cType, | |||
| int | step | |||
| ) |
A variety of stat summaries in a selected spatial window (debug-related).
This determines maximum, minimum, sum, average, either cumulatively over iterations or not cumulatively. This Model.outList command has not been used in ELM, and has not been verified for accuracy.
- Parameters:
-
fValue Model variable (local) array data label variable name nComp number of comps cType The combination (stats) type step The current iteration number
Definition at line 1050 of file Serial.c.
References ctable, getCombineIndex(), kAVE, kAVECUM, kMAX, kMAXMIN, kMIN, kSUM, kSUMCUM, msgStr, and WriteMsg().
Referenced by calc_maxmin(), and print_loc_ave().
01051 { 01052 int print, cIndex, i, cum; 01053 static int type = 11; 01054 char tmpStr[50]; 01055 if( ++type > 99) type = 11; 01056 01057 cum = ((cType == kAVECUM) || (cType == kSUMCUM)) ? 1 : 0; 01058 if(cum) { cIndex = getCombineIndex(label,step,cType,&print); } 01059 01060 switch(cType) { 01061 case kMAXMIN: 01062 sprintf(msgStr,"\nMAXMIN(%d) for %s:",step,label); 01063 for(i=0; i<nComp; i++) { sprintf(tmpStr," %f",fValue[i]); strcat(msgStr,tmpStr); } 01064 WriteMsg(msgStr,1); 01065 break; 01066 case kMAX: 01067 sprintf(msgStr,"\nMAX(%d) for %s:",step,label); 01068 for(i=0; i<nComp; i++) { sprintf(tmpStr," %f",fValue[i]); strcat(msgStr,tmpStr); } 01069 WriteMsg(msgStr,1); 01070 break; 01071 case kMIN: 01072 sprintf(msgStr,"\nMIN(%d) for %s:",step,label); 01073 for(i=0; i<nComp; i++) { sprintf(tmpStr," %f",fValue[i]); strcat(msgStr,tmpStr); } 01074 WriteMsg(msgStr,1); 01075 break; 01076 case kSUM: 01077 sprintf(msgStr,"\nSUM(%d) for %s:",step,label); 01078 for(i=0; i<nComp; i++) { sprintf(tmpStr," %f",fValue[i]); strcat(msgStr,tmpStr); } 01079 WriteMsg(msgStr,1); 01080 break; 01081 case kAVE: 01082 sprintf(msgStr,"\nAVE(%d) for %s:",step,label); 01083 for(i=0; i<nComp; i+=2) { sprintf(tmpStr," %f",fValue[i]/fValue[i+1]); strcat(msgStr,tmpStr); } 01084 WriteMsg(msgStr,1); 01085 break; 01086 case kSUMCUM: 01087 if( print == -1) { 01088 for(i=0; i<nComp; i++) ctable[cIndex].fvalue[i] = fValue[i]; 01089 01090 } else { 01091 for(i=0; i<nComp; i++) ctable[cIndex].fvalue[i] += fValue[i]; 01092 } 01093 if(print==1) { 01094 sprintf(msgStr,"\nSUMCUM(%d) for %s:",step,label); 01095 for(i=0; i<nComp; i++) { sprintf(tmpStr," %f",ctable[cIndex].fvalue[i]); strcat(msgStr,tmpStr); } 01096 WriteMsg(msgStr,1); 01097 } 01098 break; 01099 case kAVECUM: 01100 if( print == -1) { 01101 for(i=0; i<nComp; i++) ctable[cIndex].fvalue[i] = fValue[i]; 01102 01103 } else { 01104 for(i=0; i<nComp; i++) ctable[cIndex].fvalue[i] += fValue[i]; 01105 } 01106 if(print==1) { 01107 sprintf(msgStr,"\nAVECUM(%d) for %s:",step,label); 01108 for(i=0; i<nComp; i+=2) { sprintf(tmpStr," %f",ctable[cIndex].fvalue[i]/ctable[cIndex].fvalue[i+1]); strcat(msgStr,tmpStr); } 01109 WriteMsg(msgStr,1); 01110 } 01111 break; 01112 } 01113 }

| int getCombineIndex | ( | char * | name, | |
| int | step, | |||
| int | type, | |||
| int * | last | |||
| ) |
A variety of stat summaries in a selected spatial window (debug-related).
This determines maximum, minimum, sum, average, either cumulatively over iterations or not cumulatively. This Model.outList command has not been used in ELM, and has not been verified for accuracy.
- Parameters:
-
name variable name step The current iteration number type The combination (stats) type last
- Returns:
- index of the count
Definition at line 1127 of file Serial.c.
References ctable, debug, dFile, combo_table::free, max_combos_open, MAXCOMBOS, msgStr, combo_table::step, combo_table::type, and usrErr().
Referenced by Combine().
01128 { 01129 int count, index = -1; 01130 *last = 0; 01131 01132 for(count = 0; count < max_combos_open; count++) { 01133 01134 if( ctable[count].free ) { /* Determine if the slot is in use. */ 01135 if(index == -1) index = count; /* Nope, keep track of first free global stream table slot. */ 01136 if(debug) { fprintf(dFile,"cnt = %d, index = %d, free combo stream\n",count,index); fflush(dFile); } 01137 } 01138 else if( strncmp(name,ctable[count].name,25) == 0 && ctable[count].type == type ) { 01139 *last = 1; 01140 if(debug) { 01141 fprintf(dFile,"\n(%s)combo cnt = %d, step = %d, type = %d\n", 01142 name,count,step,type); 01143 fflush(dFile); 01144 } 01145 return count; 01146 } 01147 } 01148 if( index == -1 ) { 01149 index = max_combos_open; 01150 if( ++max_combos_open > MAXCOMBOS) { sprintf(msgStr,"Out of combo slots: %s, step=%d, type=%d\n",name,step,type); usrErr(msgStr); return -1; } 01151 } 01152 if(debug) { fprintf(dFile,"\n(%s)index = %d, step=%d, set-up combo\n",name,index,step); fflush(dFile); } 01153 strncpy(ctable[index].name,name,25); 01154 ctable[index].step = step; 01155 ctable[index].type = type; 01156 ctable[index].free = 0; 01157 *last = -1; 01158 return index; 01159 }

| void open_debug_outFile | ( | int | index | ) |
Open debug-related output files.
The Driver0.out and Driver1.out are two output files with debug and/or standard error-warning information. They are always produced, but have varying levels of detail (esp. Driver1.out) depending on the user-selected (at run-time) level of the "debug" variable. Driver0.out basically provides initial, static information on data and setup of the model. Driver1.out provides dynamic information on the simulation. In sensitivity analysis mode, subsequent Driver2.out, Driver3.out, etc provide dynamic information on each new simulation.
- Parameters:
-
index Index to indicate file name w/ "0", "1", and subsequent runs under a sensitivity analysis
Definition at line 1172 of file Serial.c.
References Driver_outfile, OutputPath, and ProjName.
Referenced by get_parmf(), and main().
01173 { 01174 char filename[120]; 01175 01176 sprintf(filename,"%s/%s/Output/Debug/Driver%d.out",OutputPath,ProjName,index); 01177 01178 Driver_outfile = fopen(filename,"w"); 01179 if(Driver_outfile == NULL) { 01180 fprintf(stderr,"Error, unable to open %s file.\n",filename); 01181 fprintf(stderr,"OutputPath: %s\n",OutputPath); 01182 fprintf(stderr,"Project: %s\n",ProjName); 01183 exit(0); 01184 } 01185 01186 }
| int init_config_file | ( | FILE * | vpFile, | |
| char | term1, | |||
| char | term2, | |||
| char | term3, | |||
| char | term4 | |||
| ) |
Get format definition of output configuration file (does nothing effective!).
Identifies the leading characters in the Model.outlist output configuration file that define its format, which will never change from it's value of "#2". (This is another SME component of past flexibility.)
- Parameters:
-
vpFile Pointer to the (model outlist) configuration file term1 The character "#" term2 The character "*" term3 The character "@" term4 The character "~"
- Returns:
- The (integer) format of the configuration file
Definition at line 1199 of file Serial.c.
References gTerm, and skip_white().
Referenced by readViewParms().
01200 { 01201 char test; 01202 int format,size=-1; 01203 01204 gTerm[0] = term1; 01205 gTerm[1] = term2; 01206 gTerm[2] = term3; 01207 gTerm[3] = term4; 01208 skip_white(vpFile); 01209 test = fgetc(vpFile); 01210 if(test == gTerm[0]) 01211 fscanf(vpFile,"%d",&format); 01212 else 01213 format = -1; 01214 return format; 01215 }

| int skip_cnfg_space | ( | FILE * | vpFile, | |
| char * | tch | |||
| ) |
Skip spaces in reading (model outlist) configuration file.
- Parameters:
-
vpFile Pointer to file being read tch A space
- Returns:
- negative or success
Definition at line 1223 of file Serial.c.
References cnfgFile, and gTerm.
Referenced by parse_packet().
01224 { 01225 char ch; int rv = 1; 01226 ch = *tch; 01227 while( isspace(ch) ) { 01228 if( (ch=fgetc(vpFile)) == EOF || ch == gTerm[3] || ch == '\0' ) { 01229 fclose(vpFile); cnfgFile = NULL; return -3; 01230 } 01231 } 01232 *tch = ch; 01233 return rv; 01234 }
| int parse_packet | ( | FILE * | vpFile, | |
| int * | nArgs, | |||
| char * | test | |||
| ) |
Parse through a configuration fileline to obtain the configuration commands.
Populates gCArg[][] with command info. In very early versions of ELM, this was used in reading the configuration file that translated "Stella" model equations into spatial (SME) code. Thus, much of the detail here is unused (ELM v1.0 and later) when reading the output (e.g., Model.outList) configuration file (both w/ similar syntax).
- Parameters:
-
vpFile Pointer to the (model outlist) configuration file nArgs Number of arguments in command test A character that will be used to indicate an array (in outlist, always is array, char="*")
- Returns:
- The index (aka index sequence of variables) of the ViewParm
Definition at line 1248 of file Serial.c.
References cTemp, gCArg, gTerm, kCArgDepth, kCArgWidth, and skip_cnfg_space().
Referenced by readViewParms().
01249 { 01250 static int gVPindex = -1; 01251 static char gCnfg; 01252 01253 char ch = ' ', eChar = ' '; 01254 int btype=0, argc=2, go=1, i=0, j=0, new_name = 0; 01255 01256 if(vpFile == NULL) return -3; 01257 while (1) { 01258 if ( skip_cnfg_space(vpFile, &ch) < 0) return -3; j=0; 01259 if( ch == gTerm[1] || ch == gTerm[2] ) { 01260 gCnfg = ch; ch = ' '; 01261 if ( skip_cnfg_space(vpFile, &ch) < 0 ) return -3; 01262 while ( !isspace(ch) ) { cTemp[j++] = ch; ch=fgetc(vpFile); } 01263 cTemp[j++] = '\0'; new_name = 1; 01264 gVPindex++; 01265 } 01266 else break; 01267 } 01268 strcpy(gCArg[0],cTemp); 01269 *test = gCnfg; 01270 while ( isalnum( ch ) ) { gCArg[1][i++]=ch; ch=fgetc(vpFile); } 01271 gCArg[1][i]='\0'; i=0; 01272 01273 while( 1 ) { 01274 if( ch == '(' ) { eChar = ')'; break;} 01275 else if( ch == '[' ) { eChar = ']'; break;} 01276 else if( ch == '{' ) { eChar = '}'; break;} 01277 else if( !isspace(ch) ) { return -3;} 01278 ch=fgetc(vpFile); 01279 } 01280 while (go) { 01281 while( ch=fgetc(vpFile) ) { 01282 if( ch == ',' ) { argc++; break; } 01283 if( ch == ')' || ch == '}' || ch == ']') { 01284 if( ch == eChar ) { argc++; go=0; break;} 01285 else { printf( "\nWarning: Syntax error in parse_config, var: %s\n",gCArg[0]); return -2; } 01286 } 01287 gCArg[argc][i++]=ch; 01288 if( i==(kCArgWidth-1) ) break; 01289 } 01290 if(i==0) { printf( "\nWarning: Syntax error in parse_config, var: %s\n",gCArg[0]); return -2; } 01291 else { gCArg[argc-1][i]='\0'; i=0; } 01292 if(argc == kCArgDepth) { 01293 while( ch != eChar ) ch=fgetc(vpFile); 01294 go = 0; 01295 } 01296 } 01297 *nArgs = argc; 01298 if(argc==0) { printf( "\nWarning: Syntax error in parse_config, var: %s\n",gCArg[0]); return -2; } 01299 else return gVPindex; 01300 }

| int get_number | ( | FILE * | infile | ) |
Get a numeric digit from a file.
- Parameters:
-
infile file pointer
- Returns:
- The digit
Definition at line 1308 of file Serial.c.
Referenced by goto_index().
01308 { 01309 char ch; 01310 int rv; 01311 01312 ch = fgetc(infile); 01313 if( !isdigit( ch ) ) return(-1); 01314 rv = ch - '0'; 01315 01316 while( isdigit( ch = fgetc(infile) ) ) 01317 rv = rv*10 + ( ch - '0' ); 01318 01319 return rv; 01320 }
| int goto_index | ( | FILE * | infile, | |
| char | tchar, | |||
| int | index | |||
| ) |
Go to a particular indexed/demarked point in the habitat-specfic parameter file.
- Parameters:
-
infile file pointer tchar special character that denotes either habitat or module/sector index index that points to either habitat or module/sector number
- Returns:
- succes/fail
Definition at line 1329 of file Serial.c.
References fatal(), find_char(), and get_number().
Referenced by get_hab_parm().
01330 { 01331 int rv=1, current_index=-1, itest; 01332 01333 while(current_index != index) { 01334 itest = find_char (infile, tchar); 01335 if(itest <= 0) return 0; 01336 current_index = get_number(infile); 01337 if(current_index < 0) fatal("Bad number format in dBase"); 01338 } 01339 return 1; 01340 }
| float get_Nth_parm | ( | FILE * | infile, | |
| int | pIndex, | |||
| int * | end, | |||
| int | hIndex | |||
| ) |
Get the N'th parameter in the habitat-specfic parameter file.
After finding the correct habitat and module/sector location in the file, get the value of a parameter.
- Parameters:
-
infile file pointer pIndex the index number of a parameter within a module set of parms end denotes end of file hIndex the index number of current habitat being read
- Returns:
- the parameter value
Definition at line 1353 of file Serial.c.
References debug, dFile, find_char(), msgStr, usrErr(), and WriteMsg().
Referenced by get_hab_parm().
01354 { 01355 int i=1, itest=1; 01356 float rv; char ch; 01357 01358 while ( (i++ < pIndex) && itest > 0 ) { itest = find_char ( infile, '\t' ); } 01359 if( itest == 0 ) { *end = 1; return(0.0); } 01360 01361 itest = fscanf(infile,"%f",&rv); 01362 if(itest==1) { 01363 if(debug) { 01364 sprintf(msgStr,"Habitat %d:\t%f",hIndex,rv); 01365 WriteMsg(msgStr,1); 01366 } 01367 } 01368 else if ( (ch = fgetc(infile)) == EOF ) { *end = 1; return(0.0); } 01369 else { 01370 sprintf(msgStr,"Read Error in dBase(%d)\n REad Dump:\n",itest); WriteMsg(msgStr,1); usrErr(msgStr); 01371 for(i=0; i<12; i++) { 01372 ch = fgetc(infile); 01373 if(ch==EOF) { sprintf(msgStr,"\nAt EOF\n"); WriteMsg(msgStr,1); break; } 01374 else fputc(ch,dFile); 01375 } 01376 exit(0); 01377 } 01378 *end = 0; 01379 return(rv); 01380 }
| float SMDRAND | ( | float | fminVal, | |
| float | fmaxVal | |||
| ) |
| void local_setup | ( | int | argc, | |
| char ** | argv | |||
| ) |
Does very little in serial implementation (opens a low-level debug file).
Primarily intended for parallel apps:
establish processor attributes, open a low-level debug file (in the user's current directory, set by the model execution script ("go"), and seed the pseudo-random number.
- Parameters:
-
argc number of command line arguments argv command line argument
- Returns:
- void
Definition at line 1403 of file Serial.c.
References ctable, dFile, combo_table::free, Lprocnum, max_combos_open, MAXCOMBOS, nprocs, combo_table::ocnt, procnum, recpnum, seed, tramType, and usrErr().
Referenced by main().
01404 { 01405 int i; 01406 char debugfileName[300]; 01407 nprocs[0] = 1; 01408 nprocs[1] = 1; 01409 tramType = 0; 01410 procnum = 1; 01411 recpnum[0] = 0; 01412 recpnum[1] = 0; 01413 Lprocnum = 1; 01414 /* the combo_table (ctable) used in Combine operation for spatial array summaries (debug-related) */ 01415 max_combos_open = 0; 01416 for(i = 0; i < MAXCOMBOS; i++) { 01417 ctable[i].ocnt = 0; 01418 ctable[i].free = 1; 01419 } 01420 01421 /* this debug output file is not used extensively in current code - it is opened before we get 01422 environment vars, thus is written to user's current directory ($ModelPath/$ProjName/Load if 01423 ELM is executed from the "go" script) */ 01424 dFile = fopen("ELM.debug","w"); 01425 if(dFile == NULL) { 01426 usrErr("Can't open ELM.debug file."); 01427 exit(0); 01428 } 01429 /* there are no random processes in current ELM (v2.4) w/o fire */ 01430 srand(seed); 01431 fprintf(dFile," RAND_MAX = %d\n",(int)RAND_MAX); 01432 }

| void exparam | ( | struct nodenv * | envInfo | ) |
Parallel code: effectively unused in serial implementation.
Definition at line 1439 of file Serial.c.
References nodenv::groupid, nodenv::nprocs, nodenv::procnum, and nodenv::taskid.
Referenced by setup_platform().
01440 { 01441 envInfo->procnum = 1; 01442 envInfo->nprocs = 1; 01443 envInfo->groupid = 0; 01444 envInfo->taskid = 0; 01445 01446 }
| int exgridinit | ( | int | dim, | |
| int * | nprocs | |||
| ) |
Parallel code: does nothing in serial implementation).
Definition at line 1449 of file Serial.c.
Referenced by setup_platform().
| void exgridsplit | ( | int | nprocs, | |
| int | ndim, | |||
| int | nprocs2[2] | |||
| ) |
Parallel code: effectively unused in serial implementation.
Definition at line 1453 of file Serial.c.
Referenced by setup_platform().
| void exgridcoord | ( | int | pnum, | |
| int | rnum[2] | |||
| ) |
| void exgridsize | ( | int | pnum, | |
| int | gsize[2], | |||
| int | lsize[2], | |||
| int | lstart[2] | |||
| ) |
Parallel code: effectively unused in serial implementation.
Definition at line 1467 of file Serial.c.
References nprocs, and recpnum.
Referenced by setup_grid().
01468 { 01469 int rem[2], i, j; 01470 for( i=0; i<2; i++ ) { 01471 lsize[i] = gsize[i]/nprocs[i]; 01472 rem[i] = gsize[i] - lsize[i]*nprocs[i]; 01473 if(recpnum[i]<rem[i]) lsize[i]++; 01474 for(j=0; j<recpnum[i]; j++) if( j<rem[i] && recpnum[i] >= rem[i] ) lstart[i] += (lsize[i]+1); else lstart[i] += lsize[i]; 01475 } 01476 }
| void set_async_mode | ( | FILE * | file | ) |
| void fmulti | ( | FILE * | file | ) |
| void fsingl | ( | FILE * | file | ) |
| void fasync | ( | FILE * | file | ) |
| void exchange_borders | ( | UCHAR * | map, | |
| int | size | |||
| ) |
Parallel code: does nothing in serial implementation).
Definition at line 1496 of file Serial.c.
Referenced by link_edges().
| int on_this_proc | ( | int | x, | |
| int | y | |||
| ) |
Parallel code: does nothing in serial implementation).
Definition at line 1500 of file Serial.c.
Referenced by quick_look().
| void Cplot | ( | VOIDP | Map, | |
| unsigned char | Mtype, | |||
| float | max_value, | |||
| float | min_value | |||
| ) |
| void broadcastMsg | ( | UCHAR * | msgPtr | ) |
| void broadcastInt | ( | int * | iValPtr | ) |
Parallel code: does nothing in serial implementation).
Definition at line 1512 of file Serial.c.
Referenced by get_parmf(), PTSL_ReadLists(), and read_map_dims().
| void broadcastChar | ( | UCHAR * | cPtr | ) |
| void broadcastData | ( | void * | dataPtr, | |
| int * | dataSize | |||
| ) |
| void sync_processors | ( | ) |
| void broadcastFloat | ( | void * | dataPtr | ) |
 1.5.6
1.5.6