success.h File Reference
Header file for habitat succession. More...
#include "globals.h"
Go to the source code of this file.
Data Structures | |
| struct | HabData |
| struct | Habitat |
Defines | |
| #define | MAX_SW 9999 |
| #define | AV_PER 7 |
| #define | SW_TIME_TH_W 4 |
| #define | SW_TIME_TH_N 4 |
| #define | SW_TIME_TH_S 4 |
Functions | |
| void | HabSwitch_Init (void) |
| Transfers habitat-specific parameters into data struct, allocate memory. | |
| void | alloc_hab_hist (void) |
| Allocate memory for habitat history info. | |
| unsigned char | HabSwitch (int ix, int iy, float *Water, float *Nutrient, int *Fire, float *Salinity, unsigned char *HAB) |
| Switches habitats (the option that is currently used). | |
| int | InHab (float Var, struct HabData Params) |
| Defines the suitability of current habitat relative to existing conditions. | |
| void | init_pvar (VOIDP Map, UCHAR *mask, unsigned char Mtype, float iv) |
| Initialize a variable to a value. | |
| VOIDP | nalloc (unsigned mem_size, const char var_name[]) |
| Allocate memory for a variable. | |
Variables | |
| unsigned long * | HabHist |
| struct Habitat | Habi [MAX_NHAB] |
| float * | HP_SfDepthLo |
| float * | HP_SfDepthHi |
| float * | HP_SfDepthInt |
| float * | HP_PhosLo |
| float * | HP_PhosHi |
| float * | HP_PhosInt |
| float * | HP_FireInt |
| float * | HP_SalinLo |
| float * | HP_SalinHi |
| float * | HP_SalinInt |
| int | habNumTot |
Detailed Description
Header file for habitat succession.
This defines or declares variables & functions that are global to Success.c.
Note: documented with Doxygen, which expects specific syntax within special comments.
The Everglades Landscape Model (ELM).
last updated: Sep 2011
Definition in file success.h.
Define Documentation
| #define MAX_SW 9999 |
| #define AV_PER 7 |
Number of days that constitutes the period of averaging - use weekly (7 d); should be <100
Definition at line 25 of file success.h.
Referenced by HabSwitch().
| #define SW_TIME_TH_W 4 |
Switching time threshold for water - use 4 d for weekly AV_PER (i.e., majority of week)
Definition at line 26 of file success.h.
Referenced by HabSwitch().
| #define SW_TIME_TH_N 4 |
Switching time threshold for nutrients - use 4 d for weekly AV_PER (i.e., majority of week)
Definition at line 27 of file success.h.
Referenced by HabSwitch().
| #define SW_TIME_TH_S 4 |
Switching time threshold for salinity - use 4 d for weekly AV_PER (i.e., majority of week)
Definition at line 28 of file success.h.
Referenced by HabSwitch().
Function Documentation
| void HabSwitch_Init | ( | void | ) |
Transfers habitat-specific parameters into data struct, allocate memory.
Definition at line 25 of file Success.c.
References alloc_hab_hist(), Habi, habNumTot, HP_FireInt, HP_PhosHi, HP_PhosInt, HP_PhosLo, HP_SalinHi, HP_SalinInt, HP_SalinLo, HP_SfDepthHi, HP_SfDepthInt, HP_SfDepthLo, HabData::Lhi, HabData::Llo, msgStr, Habitat::Nutrient, Habitat::PFin, HabData::Pin, Habitat::Salinity, usrErr(), Habitat::Water, and WriteMsg().
00026 { 00027 int ii; 00028 00029 /* put these habitat-specific parameters into a concise data struct for use here */ 00030 for( ii = 0; ii < habNumTot; ii++) { 00031 Habi[ii].Water.Llo = HP_SfDepthLo[ii+1]; 00032 Habi[ii].Water.Lhi = HP_SfDepthHi[ii+1]; 00033 Habi[ii].Water.Pin = HP_SfDepthInt[ii+1]; 00034 Habi[ii].Nutrient.Llo = HP_PhosLo[ii+1]; 00035 Habi[ii].Nutrient.Lhi = HP_PhosHi[ii+1]; 00036 Habi[ii].Nutrient.Pin = HP_PhosInt[ii+1]; 00037 Habi[ii].PFin = HP_FireInt[ii+1]; 00038 Habi[ii].Salinity.Llo = HP_SalinLo[ii+1]; 00039 Habi[ii].Salinity.Lhi = HP_SalinHi[ii+1]; 00040 Habi[ii].Salinity.Pin = HP_SalinInt[ii+1]; 00041 } 00042 00043 alloc_hab_hist( ); 00044 sprintf (msgStr, "Succession ON, module OK for %d habitats...", ii ); usrErr(msgStr); WriteMsg(msgStr,1); 00045 00046 return; 00047 }
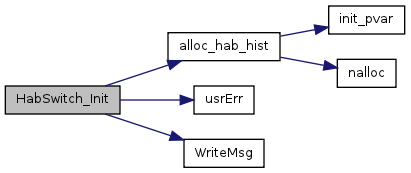
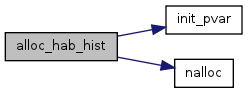
| void alloc_hab_hist | ( | void | ) |
Allocate memory for habitat history info.
Definition at line 137 of file Success.c.
References HabHist, habNumTot, init_pvar(), nalloc(), s0, and s1.
Referenced by HabSwitch_Init().
00138 { 00139 HabHist = (unsigned long *) nalloc(sizeof(unsigned long)*(s0+2)*(s1+2)*(habNumTot),"HabHist"); 00140 init_pvar(HabHist,NULL,'i',0); 00141 }
| unsigned char HabSwitch | ( | int | ix, | |
| int | iy, | |||
| float * | Water, | |||
| float * | Nutrient, | |||
| int * | Fire, | |||
| float * | Salinity, | |||
| unsigned char * | HAB | |||
| ) |
Switches habitats (the option that is currently used).
Returns the number of habitat to switch to based on the length of the weekly (or other) averaged period of favorable conditions. In this case the period of habitat favorable conditions is calculated as the number of weeks (or other time intervals) during which the habitat conditions matched the specified for a period longer than the given one - SW_TIME_TH. The switching occurs as soon as the number of such successfull weeks exceeds the time specified in Pin
- Parameters:
-
ix Model domain row iy Model domain column Water Current water depth data array Nutrient Current nutrient conc. data array Fire Fire data array (unused, no fire implemented) Salinity Current salinity conc. data array HAB Habitat-type data array
- Returns:
- habitat type of cell
Definition at line 66 of file Success.c.
References AV_PER, cell, conv_kgTOmg, HabHist, Habi, habNumTot, InHab(), MAX_NHAB, Habitat::Nutrient, HabData::Pin, Habitat::Salinity, SimTime, SW_TIME_TH_N, SW_TIME_TH_S, SW_TIME_TH_W, T, and simTime::TIME.
00067 { 00068 int StateW, StateN, StateS, i; 00069 int HabW[MAX_NHAB], HabN[MAX_NHAB], HabS[MAX_NHAB], DW, DN, DS; 00070 int cell = T(ix,iy); 00071 int hab = HAB[cell]; /* current habitat in the cell */ 00072 00073 /* define the habitat type that matches the existing water conditions */ 00074 00075 /* HabHist is an array of integers : ssSnnNwwW, where 00076 ww are the two digits for the number of weeks in the water level favorable conditions, 00077 W the number of days in those conditions during the current week; 00078 nn are the two digits for the number of weeks being in the nutrient favorable conditions, 00079 N the number of days in those conditions during the current week; 00080 ss are the two digits for the number of weeks in the salinity favorable conditions, 00081 S the number of days in those conditions during the current week; 00082 00083 The switching occurs when all of these histories exceed the habitat specific 00084 periods Pin. 00085 */ 00086 for ( i = 0; i < habNumTot; i++ ) 00087 { 00088 DW = HabHist[cell*habNumTot +i]%10; 00089 HabW[i] = (HabHist[cell*habNumTot +i]/10)%100; 00090 DN = (HabHist[cell*habNumTot +i]/1000)%10; 00091 HabN[i] = HabHist[cell*habNumTot +i]/10000; 00092 DS = (HabHist[cell*habNumTot +i]/1000000)%10; 00093 HabS[i] = HabHist[cell*habNumTot +i]/10000000; 00094 00095 /* when the averaging time elapses, if #favorable days exceeds a threshold (using 4 for 7 day AV_PER), increment the period (weeks) history */ 00096 if ((int)SimTime.TIME%AV_PER == 0) 00097 { 00098 if (DW > SW_TIME_TH_W) HabHist[cell*habNumTot +i] = HabHist[cell*habNumTot +i] - DW + 100; 00099 else HabHist[cell*habNumTot +i] = 0; 00100 if (DN > SW_TIME_TH_N) HabHist[cell*habNumTot +i] = HabHist[cell*habNumTot +i] - DN*1000 + 10000; 00101 else HabHist[cell*habNumTot +i] = 0; 00102 if (DS > SW_TIME_TH_S) HabHist[cell*habNumTot +i] = HabHist[cell*habNumTot +i] - DS*1000000 + 10000000; 00103 else HabHist[cell*habNumTot +i] = 0; 00104 } 00105 00106 /* check what habitat type the existing conditions match; increment the day# if conditions are favorable */ 00107 if ( InHab (Water[cell], Habi[i].Water) ) HabHist[cell*habNumTot + i] = HabHist[cell*habNumTot + i] + 1; 00108 if ( InHab (Nutrient[cell]*conv_kgTOmg, Habi[i].Nutrient) ) HabHist[cell*habNumTot + i] = HabHist[cell*habNumTot + i] + 1000; 00109 if ( InHab (Salinity[cell], Habi[i].Salinity) ) HabHist[cell*habNumTot + i] = HabHist[cell*habNumTot + i] + 1000000; 00110 } 00111 00112 /* check if the historical conditions for switching hold */ 00113 for ( i = 0; i < habNumTot; i++ ) 00114 if ( HabW[i] >= Habi[i].Water.Pin && HabN[i] >= Habi[i].Nutrient.Pin && HabS[i] >= Habi[i].Salinity.Pin ) 00115 { HabHist[cell*habNumTot +i] = 0; 00116 return (i+1); /* returns new hab, HAB[] is offset by 1 from these array IDs */ 00117 } 00118 00119 return hab; 00120 }

| int InHab | ( | float | Var, | |
| struct HabData | Params | |||
| ) |
Defines the suitability of current habitat relative to existing conditions.
- Parameters:
-
Var The model variable being considered Params Struct of habitat switching parameters
- Returns:
- true/false
Definition at line 128 of file Success.c.
References HabData::Llo, and MAX_SW.
Referenced by HabSwitch().
00129 { 00130 if ( (Var <= Params.Lhi || Params.Lhi >= MAX_SW) && Var >= Params.Llo ) 00131 return 1; 00132 else return 0; 00133 }
| void init_pvar | ( | VOIDP | Map, | |
| UCHAR * | mask, | |||
| unsigned char | Mtype, | |||
| float | iv | |||
| ) |
Initialize a variable to a value.
- Parameters:
-
Map array of data mask data mask Mtype the data type of the map data iv the value used to initialize variable
Definition at line 1008 of file Driver_Utilities.c.
References numBasn, s0, s1, and T.
01009 { 01010 int i0, i1; 01011 01012 switch(Mtype) { 01013 case 'b' : /* added double for (non-map) basin (b) array budget calcs */ 01014 for(i0=0; i0<=numBasn; i0++) { 01015 ((double*)Map)[i0] = iv; 01016 } 01017 break; 01018 case 'l' : /* added double (l == letter "ell" ) for map arrays */ 01019 for(i0=0; i0<=s0+1; i0++) 01020 for(i1=0; i1<=s1+1; i1++) { 01021 if(mask==NULL) ((double*)Map)[T(i0,i1)] = iv; 01022 else if ( mask[T(i0,i1)] == 0 ) ((double*)Map)[T(i0,i1)] = 0; 01023 else ((double*)Map)[T(i0,i1)] = iv; 01024 } 01025 break; 01026 case 'f' : 01027 for(i0=0; i0<=s0+1; i0++) 01028 for(i1=0; i1<=s1+1; i1++) { 01029 if(mask==NULL) ((float*)Map)[T(i0,i1)] = iv; 01030 else if ( mask[T(i0,i1)] == 0 ) ((float*)Map)[T(i0,i1)] = 0; 01031 else ((float*)Map)[T(i0,i1)] = iv; 01032 } 01033 break; 01034 case 'i' : case 'd' : 01035 for(i0=0; i0<=s0+1; i0++) 01036 for(i1=0; i1<=s1+1; i1++) { 01037 if(mask==NULL) ((int*)Map)[T(i0,i1)] = (int)iv; 01038 else if ( mask[T(i0,i1)] == 0 ) ((int*)Map)[T(i0,i1)] = 0; 01039 else ((int*)Map)[T(i0,i1)] = (int)iv; 01040 } 01041 break; 01042 case 'c' : 01043 for(i0=0; i0<=s0+1; i0++) 01044 for(i1=0; i1<=s1+1; i1++) { 01045 if(mask==NULL) ((unsigned char*)Map)[T(i0,i1)] = (unsigned char) '\0' + (int) iv; 01046 else if ( mask[T(i0,i1)] == 0 ) ((unsigned char*)Map)[T(i0,i1)] = '\0'; 01047 else ((unsigned char*)Map)[T(i0,i1)] = (unsigned char) '\0' + (int) iv; 01048 } 01049 break; 01050 01051 default : 01052 printf(" in default case\n"); 01053 break; 01054 } 01055 }
| VOIDP nalloc | ( | unsigned | mem_size, | |
| const char | var_name[] | |||
| ) |
Allocate memory for a variable.
- Parameters:
-
mem_size The size of memory space var_name The variable's name
- Returns:
- Pointer to that memory
Definition at line 1857 of file Driver_Utilities.c.
References Exit(), fasync(), fmulti(), total_memory, and VOIDP.
01858 { 01859 VOIDP rp; 01860 01861 01862 if(mem_size == 0) return(NULL); 01863 rp = (VOIDP)malloc( mem_size ); 01864 total_memory += mem_size; 01865 fasync(stderr); 01866 if( rp == NULL ) { 01867 fprintf(stderr,"Sorry, out of memory(%d): %s\n",mem_size,var_name); 01868 Exit(0); 01869 } 01870 fmulti(stderr); 01871 return(rp); 01872 }
Variable Documentation
| unsigned long* HabHist |
HabHist is an array of integers : nnNssS, where nn are the two digits for the number of weeks being in the nutrient favorable conditions, N the number of days in those conditions during the current week; similarly ss are the two digits for the number of weeks in the water level favorable conditions and S is the number of days during the current week in that state. The switching occurs when both of these histories exceed the habitat specific periods Pin.
Definition at line 37 of file success.h.
Referenced by alloc_hab_hist(), and HabSwitch().
| float* HP_SfDepthLo |
Habitat-specific parameter. _Units_: m; _Brief_: Lower Depth tolerance for Surface Water Depth
Definition at line 73 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_SfDepthHi |
Habitat-specific parameter. _Units_: m; _Brief_: Higher Depth tolerance for Surface Water Depth
Definition at line 74 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_SfDepthInt |
Habitat-specific parameter. _Units_: days; _Brief_: Time Interval for staying within Surface Water Depth range
Definition at line 75 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_PhosLo |
Habitat-specific parameter. _Units_: mgP/kg soil; _Brief_: Lower concentration tolerance for soil total Phosphorus
Definition at line 76 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_PhosHi |
Habitat-specific parameter. _Units_: mgP/kg soil; _Brief_: Higher concentration tolerance for soil total Phosphorus
Definition at line 77 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_PhosInt |
Habitat-specific parameter. _Units_: days; _Brief_: Time Interval for staying within soil total Phosphorus range
Definition at line 78 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_FireInt |
Habitat-specific parameter. _Units_: days; _Brief_: UNUSED. Time Interval since last Fire
Definition at line 79 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_SalinLo |
Habitat-specific parameter. _Units_: g/L; _Brief_: Lower concentration tolerance for soil water salinity
Definition at line 80 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_SalinHi |
Habitat-specific parameter. _Units_: g/L; _Brief_: Higher concentration tolerance for soil water salinity
Definition at line 81 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| float* HP_SalinInt |
Habitat-specific parameter. _Units_: days; _Brief_: Time Interval for staying within soil water salinity range
Definition at line 82 of file unitmod_habparms.h.
Referenced by alloc_memory(), HabSwitch_Init(), and ReadHabParms().
| int habNumTot |
total number of habitat-types used in model
Definition at line 49 of file serial.h.
Referenced by alloc_hab_hist(), get_hab_parm(), HabSwitch(), and HabSwitch_Init().
 1.5.6
1.5.6